Abstract
Homologous recombination provides high-fidelity DNA repair in all domains of life. Live-cell fluorescence microscopy offers the opportunity to image individual recombination events in real-time, providing insights into the in vivo biochemistry of the proteins and DNA molecules involved, as well as the cellular organization of the homologous recombination process. . Here we review the cellular biological aspects of mitotic homologous recombination with a focus on Saccharomyces cerevisiae and mammalian cells, but we will also draw on findings from other experimental systems.
Key topics of this review include the stoichiometry and dynamics of recombination complexes in vivo, the choreography of assembly and disassembly of recombination proteins at sites of DNA damage, the mobilization of damaged DNA during homology search, and the functional compartmentalization of the nucleus with respect to The ability of homologous recomeniotic recombination is initiated by double-stranded DNA breaks Extensive base-pair interactions cannot occur between two intact DNA double helices.
Therefore, the DNA synapse that is critical for general recombination in meiosis can begin only after one DNA strand of one DNA helix has been exposed and its nucleotides have become available for pairing with another DNA helix. In the absence of direct experimental evidence, theoretical models were proposed based on the idea that a break needed to be made in only one of the two strands of a DNA helix to produce the exposed DNA strand required for DNA synapsis.
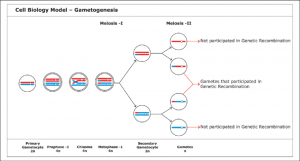
This break in the phosphodiester backbone was thought to allow one end of the nicked strand to separate from its associated base-paired strand, freeing it to form a short heteroduplex with an intact second DNA helix, thus beginning synapsis. Models of this type are reasonable in theory and have been described in textbooks for nearly 30 years.
In the early 1990s, sensitive biochemical techniques became available to determine the actual structure of recombination intermediates that are formed on yeast chromosomes at various stages of meiosis. These studies revealed that general recombination is initiated by a special endonuclease that simultaneously cuts both strands of the double helix, creating a complete break in the DNA molecule.
The 5′ ends at the break are then chewed up by an exonuclease, creating protruding single-stranded 3′ ends. It is these individual strands that seek out a homologous DNA helix to pair with, leading to the formation of a joint molecule between a maternal and paternal chromosomal pairing.
Recombinant ligand applications in molecular and cell biology
1. Cell proteomics
Proteome analysis relies on the availability of high-quality binding reagents. They are required for molecular analysis of specific target proteins and their spatiotemporal cellular abundance using standard detection methods such as microscopy and Western blot analysis. Furthermore, binder-mediated affinity purification in combination with systems biology techniques has contributed to a comprehensive understanding of cellular interactomes in development and disease.
Although such experiments have historically been performed with conventional antibodies, the use of recombinant binding reagents for proteomic applications is becoming increasingly popular, and international consortia have been set up with the long-term goal of covering the entire human proteome with renewable binders (Taussig et al., 2007; Colwill and Gräslund, 2011).
Biochemical and proteomic analyzes of crude samples include affinity-based purification or depletion of specific components using chromatography columns, however, requiring large amounts of affinity material. Therefore, expensive monoclonal and polyclonal antibodies are less attractive for purifying or depleting endogenous proteins, while inexpensive immobilized metal affinity chromatography (IMAC) systems, such as the Ni-NTA (nitrilotriacetic acid)/His- tag, are widely used but also restricted to artificially tagged proteins. Consequently, recombinant binders that can be produced cheaply in large quantities offer valuable alternatives for immunoaffinity chromatography of biological samples.
2. Confirmation and PTM sensors
As discussed in the structural biology section, affinity reagents can detect and stabilize specific conformational states of proteins. Consequently, reformulating such binders as intracellular biosensors potentially allows spatiotemporal analysis of specific conformational changes and their biological implications in living Cell Biology Recombinants. Such an application was shown for a nanobody that binds to the activated conformation of the β2-adrenoceptor (Rasmussen et al., 2011a).
Fusion of this binder with GFP informs the subcellular localization of activated β2-adrenoceptor and leads to the identification of endosomal membranes as initiation sites for acute G protein-coupled receptor signalling, previously thought to occur exclusively at the membrane. plasma (Irannejad et al., 2012). al., 2013). Similarly, a DARPin that recognizes the activated phosphorylated conformation of the extracellular signal-regulated kinase (ERK; Kummer et al., 2012) was conjugated with a solvatochromic merocyanine to quantitatively report active localization of pERK in living cells in the absence and presence of an inhibitor of the upstream regulatory kinase MEK1/2 (Kummer et al., 2013).
Interestingly, this DARPin recognizes a conformational change within the activation cycle that is dependent on activation and thus indirectly informs the main modification, ERK phosphorylation. Like conventional antibodies, recombinant binders can be raised against PTMs and non-protein epitopes. Therefore, the application of recombinant binders in living cells paves the way towards dynamic PTM analysis, as demonstrated with a GFP-tagged scFv that specifically recognizes histone acetylation (H3K9ac) and is used for dynamic chromatin monitoring. epigenetics in living cells and living animals. modulation (Sato et al., 2013).
3. PPI
Screening for PPIs is often done in vitro or in yeast. In living cells, PPIs can be tested with FRET, which is technically demanding, or with protein fragment complementation assays, which do not report in real-time and are irreversible. In contrast, the newly developed fluorescent-3-hybrid (F3H) assay is based on recombinant immune recruitment and allows dynamic and reversible monitoring of PPIs in living mammalian cells with a simple optical readout GFP fusions with proteins of interest (bait) were recruited into discrete subcellular structures, such as artificial Laco DNA arrays, by fusing GBP with the Lac repressor.
In addition, major natural satellite repeats, the nuclear periphery, and cytoplasmic structures such as centrosomes can be used as anchor points. The enrichment of red fluorescent prey proteins at the respective docking sites indicates a specific interaction. The nanobody-based F3H assay is also suitable for drug discovery, as small molecule compounds or peptides that prevent or disrupt PPIs can be identified. With this F3H assay, for example, disruption of p53-HDM2 interaction by potential anticancer drugs such as Nutlin-3 was monitored in live cells, providing direct information on dose-response, kinetics, and bioavailability ( Herce et al., 2013).